Tumor mutation burden: a promising predictive biomarker for cancer immunotherapy
Targeting the inhibitory programmed death-ligand 1 (PD-L1)/programmed death-1 (PD-1) axis with the immune checkpoint blocker nivolumab (Opdivo®) or atezolizumab (Tecentriq®), and the cytotoxic T-Lymphocyte Associated Protein 4 (CTLA4 or CD152) with the immune checkpoint blocker ipilimumab (Yervoy™) or tremelimumab, has become the face of immuno-oncology (1-5).
Treatment modalities with established predictive biomarkers that provide information about the effect of a therapeutic intervention and can be a target for therapy, and prognostic biomarkers that provide information about patients’ overall cancer outcome, regardless of therapy, can translate into precision medicine. In the case of predictive biomarkers, patients who will benefit from the treatment can be identified and, subsequently, targeted, leading to a wider-successfully-targeted patient populations, and a higher survival rate. Immunotherapy, in its current shape, lacks well-established predictive biomarkers, and, hence, applying immunotherapy to patients, unfortunately, remains more of a blind-not-precision-medicine. The only currently-adopted predictive biomarker, PD-L1 expression in the tumor microenvironment (6, 7), suffers several limitations (8, 9).
In melanoma, Snyder and colleagues (10) found that tumor mutation burden (TMB) measured by whole-exome sequencing (WES), was a predictor of increased survival in patients taking ipilimumab or tremelimumab. Hugo and coworkers (11) noted that higher TMB was associated with improved survival in patients with metastatic melanoma receiving PD-1 immune checkpoint blockade. Whilst, Van Allen et al. (12) found that higher TMB correlated with higher clinical benefit in patients with metastatic cutaneous melanoma taking ipilimumab.
In colorectal cancer, Le and coworkers (13) noted that mismatch-repair status, a part of TMB, predicted clinical benefit in patients taking pembrolizumab (Keytruda®, PD-1 inhibitor in lymphocytes).
In non-small cell lung cancer (NSCLC), Rizvi and colleagues (14) demonstrated that a higher somatic nonsynonymous mutation burden measured by WES, was associated with improved overall response rates (ORRs), progression-free survival (PFS) and durable clinical benefit in patients taking pembrolizumab. This observation aligned with metastatic NSCLC, where Carbone and coworkers (15), in the CheckMate 026 study, noted that a higher TMB was associated with higher response rates in patients receiving nivolumab in combination with platinum doublet chemotherapy as first-line therapy. Specifically, patients with higher TMB alongside higher PD-L1 expression, achieved best clinical outcomes. Hellman and coworkers (16), in CheckMate 012, noted that that TMB could identify advanced NSCLC patients, who would benefit from the nivolumab/ipilimumab combination as first line therapy. This spurred the conduction of CheckMate 568, where Ramalingam and colleagues (17), found that in NSCLC, a TMB score of ≥ 10 mutations/megabase set responders from non-responders apart, to the nivolumab/ipilimumab combination as first line therapy, regardless of their PD-L1 levels. These studies bonify the clinical benefit of TMB as a predictive biomarker in the setting of advanced NSCLC.
Tumors with higher TMB are perceived to have more immunogenic neoantigens that can be recognized by the immune system in response to immune checkpoint inhibitors (18, 19) (see Fig. 1 below).
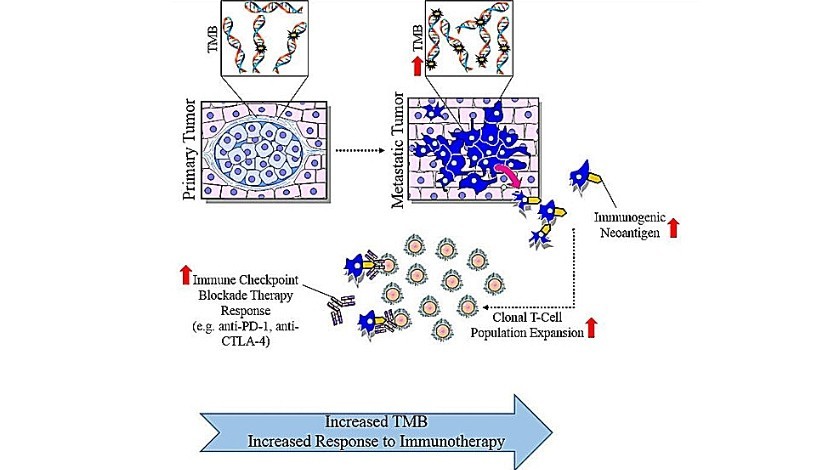
Figure 1: Medical illustration depicting tumor mutation burden
Rizvi et al. (20) noted that in advanced NSCLC, a higher TMB measured by next-generation sequencing (NGS) versus WES, correlated with durable clinical benefit and improved ORRs and PFS rates, in patients receiving immune checkpoint inhibitors. Kowanetz and coworkers (21) reported that in advanced NSCLC, a higher TMB measured by NGS, was associated with higher ORRs, PFS, and overall survival (OS), in patients taking atezolizumab, independent of their PD-L1 status.
In various cancers, Goodman and colleagues (22) found that a higher TMB measured by NGS, correlated with higher ORRs, PFS, and OS, in patients taking different immune checkpoint inhibitors. In the previous three studies (20-22), investigators defined high TMB at different thresholds and used different NGS panels. In addition, NGS is less expensive in comparison with WES in providing TMB molecular data. Two studies (20, 23) looked into TMB, assessed by NGS versus WES, in relation to clinical outcomes with immune checkpoint inhibitors, and concluded that TMB is a predictive biomarker of value. Opting for NGS in clinical practice after conducting larger clinical trials that substantiate NGS’s value over WES in assessing TMB, and synchronizing the threshold for high TMB, alongside unifying the NGS panels, are areas of improvement.
In many of the aforementioned studies, a higher TMB correlated with higher clinical benefit, irrespective of PD-L1 levels. This already sets TMB at an advantage over PD-L1 expression, as a better predictive biomarker.
Another method to assess TMB is cell-free DNA in peripheral blood, which was shown to be predictive of the benefit of immune checkpoint inhibition (24). Aside from TMB, there are other promising immunotherapy predictive biomarkers in the making (25) such as: imaging biomarkers, peripheral blood T cells (26), T-cell receptor clonality, tumor-infiltrating lymphocytes (27, 28) and immune gene signatures (29).
As promising as TMB may be (22), limitations include: cost, patient access to testing, long turnaround time, diverse testing platforms, tumor genomic heterogeneity and specimen availability.
The research and development of cost-effective, patient accessible and precise predictive biomarkers, remains a clinical priority to derive most patient-benefit from immunotherapy. TMB is one candidate worth-considering.
References:
- Pardoll DM, The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12, 252–264 (2012).
- Tang J, Yu JX, Hubbard-Lucey VM, Neftelinov ST, Hodge JP, Lin Y. Trial watch: The clinical trial landscape for PD1/PDL1 immune checkpoint inhibitors. Nat. Rev. Drug Discov. 17, 854–855 (2018).
- Ribas A, Wolchok JD, Cancer immunotherapy using checkpoint blockade. Science 359, 1350-+ (2018).
- Sharma P, Allison JP., The future of immune checkpoint therapy. Science 348, 56–61 (2015).
- Wei SC, C. Duffy R, Allison JP, Fundamental mechanisms of immune checkpoint blockade therapy. Cancer Discov. 8, 1069–1086 (2018).
- Rosenberg JE, Hoffman-Censits J, Powles Tet al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387, 1909–1920 (2016).
- Gibney GT, Weiner LM, Atkins MB, Predictive biomarkers for checkpoint inhibitor-based immunotherapy. Lancet Oncology 17, E542–E551 (2016).
- Rimm DL, Han G, Taube JM et al. A prospective, multi-institutional, pathologist-based assessment of 4 immunohistochemistry assays for PD-L1 expression in non-small cell lung cancer. JAMA Oncol. 3, 1051–1058 (2017).
- McLaughlin J, Han G, Schalper KA et al. Quantitative assessment of the heterogeneity of pd-l1 expression in non-small-cell lung cancer. JAMA Oncol. 2, 46–54 (2016).
- Snyder A, Makarov V, Merghoub T, et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
- Hugo W, Zaretsky JM, Sun L et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 168, 542 (2017).
- E. M. Van Allen, Genomic correlates of response to CTLA-4 blockade in metastatic melanoma (vol 352, aaf8264, 2016). Science 353, 229–229 (2016).
- Le DT, Uram JN, Wang H et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509-2520 (2015).
- Rizvi NA, Hellmann MD, Snyder A et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
- Carbone DP, Reck M, Paz-Ares L, et al. First-Line nivolumab in stage IV or recurrent non-small-cell lung cancer. N. Engl. J. Med. 376, 2415–2426 (2017).
- Hellmann MD, Rizvi NA, Goldman JW et al. Nivolumab plus ipilimumab as first-line treatment for advanced non-small-cell lung cancer (CheckMate 012): results of an open-label, phase 1, multicohort study. Lancet Oncol. 18, 31–41 (2017).
- Ramalingam SS, Hellmann MD, Awad MM et al. Abstract CT078: Tumor mutational burden (TMB) as a biomarker for clinical benefit from dual immune checkpoint blockade with nivolumab (nivo) + ipilimumab (ipi) in first-line (1L) non-small cell lung cancer (NSCLC): identification of TMB cutoff from CheckMate 568. Presented at: AACR Annual Meeting 2018; IL, USA, April 14–18, 2018 (2018).
- Schumacher TN, Schreiber RD, Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
- McGranahan N, Furness AJ, Rosenthal R et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
- Rizvi H, Sanchez-Vega F, La K et al. Molecular determinants of response to anti-programmed cell death (PD)-1 and anti-programmed death-ligand 1 (PD-L1) blockade in patients with non-small-cell lung cancer profiled with targeted next-generation sequencing. J. Clin. Oncol. 36, 633–641 (2018).
- Kowanetz M, Zou W, Shames DS et al. Tumor mutation load assessed by FoundationOne (FM1) is associated with improved efficacy of atezolizumab (atezo) in patients with advanced NSCLC. Ann. Oncol. 27, 15–42 (2016).
- Goodman AM, Kato S, Bazhenova L et al. Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol. Cancer Ther. 16, 2598–2608 (2017).
- Campesato LF, Barroso-Sousa R, Jimenez L et al. Comprehensive cancer-gene panels can be used to estimate mutational load and predict clinical benefit to PD-1 blockade in clinical practice. Oncotarget 6, 34221–34227 (2015).
- DR Gandara, M Kowanetz, TSK Mok et al. Blood-based biomarkers for cancer immunotherapy: tumor mutational burden in blood (bTMB) is associated with improved atezolizumab (atezo) efficacy… Ann. Oncol. 28(Suppl_5), v460–v496 (2017).
- Gibney GT, Weiner LM, Atkins MB, Predictive biomarkers for checkpoint inhibitor-based immunotherapy. Lancet Oncol. 17, e542–e551 (2016).
- Kamphorst AO, Pillai RN, Yang S et al. Proliferation of PD-1+ CD8 T cells in peripheral blood after PD-1-targeted therapy in lung cancer patients. Proc. Natl. Acad. Sci. USA 114, 4993–4998 (2017).
- Daud AI, Loo K, Pauli ML et al. Tumor immune profiling predicts response to anti-PD-1 therapy in human melanoma. J. Clin. Invest. 126, 3447–3452 (2016).
- Tumeh PC, Harview CL, Yearley JH et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
- NB Jamieson, AV Maker, Gene-expression profiling to predict responsiveness to immunotherapy. Cancer Gene Ther. 24, 134–140 (2017).
Suggested Reading:
- Related Oncology Central paper: oncology-central.com/2018/07/09/patient-selection-anti-pd-1pd-l1-therapy-advanced-non-small-cell-lung-cancer-implications-clinical-practice/
- On immune checkpoint inhibition: references 1-5
- On immunogenic neoantigens: reference 18
- On TMB as a predictive biomarker: references 20 and 22
Author profile: Balkees Abderrahman, M.D. is the Dallas/Ft. Worth Living Legend Fellow of Cancer Research at the Department of Breast Medical Oncology, the University of Texas MD Anderson Cancer Center (TX, USA) and a PhD trainee under model “Individuals of Very High Quality” at the Faculty of Biological Sciences, the University of Leeds (West Yorkshire, UK). Balkees is a regular contributor to Oncology Central; you can view her full biography here.