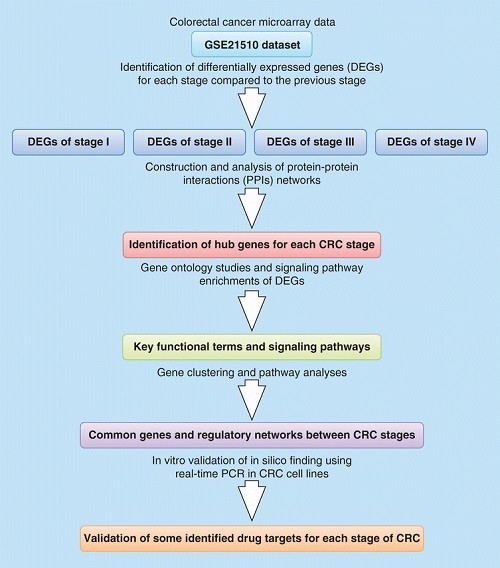
Key genes and regulatory networks involved in the initiation, progression and invasion of colorectal cancer
Abstract
Aim: Until now, identification of drug targets for treatment of patients with specific stages of colorectal cancer (CRC) has remained a challenging field of research. Herein, we aimed to identify the key genes and regulatory networks involved in each stage of CRC. Results: The results of gene expression profiles were integrated with protein–protein interaction networks, and topologically analyzed. The most important regulatory genes (e.g., CDK1, UBC, ESR1 and ATXN1) and signaling pathways (e.g., Wnt, MAPK and JAK-STAT) in CRC initiation, progression and metastasis were identified. In vitro analysis confirmed some in silico findings. Conclusion: Our study introduces functional hub genes, subnetworks, prioritizes signaling pathways and novel biomarkers in colorectal cancer that may guide further development of targeted therapy programs.
Lay abstract
A comprehensive view of the molecular mechanisms involved in colorectal cancer (CRC) pathogenesis has been demonstrated. Cell cycle and signal transduction genes were contributors to CRC initiation and progression. In contrast to stage IV, the stages I–III of CRC shared a notable number of common genes suggesting their close gene signatures. The efficacy of these predictions has been validated using two colon cancer cell lines in vitro.